Research in the Li Lab
Our research centers on studying the chemistry and biology of natural products to address major challenges in infectious disease. To combat antibiotic resistance, compounds with novel antimicrobial mechanisms are urgently needed. Natural products are highly successful as life-saving medicines, however, for many natural products, a lack of knowledge about their mode of action has severely limited their utility. We take a multidisciplinary approach to address these challenges, combining natural product chemistry, enzyme chemistry, and molecular microbiology. We elucidate the structures, in vivo mechanisms, and biosynthesis of natural-product antibiotics. In parallel, we characterize the molecular mechanisms by which pathogens infect or damage their hosts. These research areas are described below:
1. Elucidating the mode of action of natural-product antibiotics
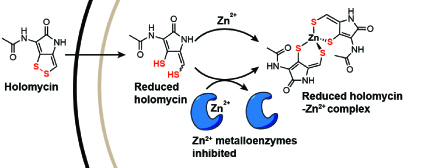 Natural products have provided many powerful antimicrobial scaffolds, such as beta-lactams and macrolides. Many natural products with potent antimicrobial activities have been overlooked in part due to a lack of understanding of their mode of action, such as the dithiolopyrrolones (DTPs). DTPs are promising antibiotics because they inhibit a broad spectrum of multidrug-resistant pathogens and exhibit low frequency of resistance. They contain a hetero-bicyclic scaffold and an unusual ene-disulfide, but it was unknown how the unique structure leads to the potent bioactivity. We elucidated the mode of action, redox chemistry, and biosynthesis of the DTP family. Our work identified metal chelation as a promising strategy to overcome antibiotic resistance. Our findings also set the stage for exploring DTPs as antimicrobials and antibiotic adjuvants against multidrug-resistant infections.
Natural products have provided many powerful antimicrobial scaffolds, such as beta-lactams and macrolides. Many natural products with potent antimicrobial activities have been overlooked in part due to a lack of understanding of their mode of action, such as the dithiolopyrrolones (DTPs). DTPs are promising antibiotics because they inhibit a broad spectrum of multidrug-resistant pathogens and exhibit low frequency of resistance. They contain a hetero-bicyclic scaffold and an unusual ene-disulfide, but it was unknown how the unique structure leads to the potent bioactivity. We elucidated the mode of action, redox chemistry, and biosynthesis of the DTP family. Our work identified metal chelation as a promising strategy to overcome antibiotic resistance. Our findings also set the stage for exploring DTPs as antimicrobials and antibiotic adjuvants against multidrug-resistant infections.
2. Identifying pathogen-derived natural-product virulence factors
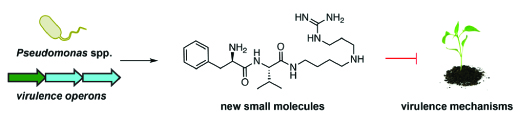
We aim to understand the roles of pathogen-derived natural products that enable pathogens to infect their hosts (virulence factors). Understanding how pathogens impair host defenses is essential to combating infectious diseases. Pathogens secrete both proteins and small molecules to damage their hosts, but the role of small molecules in virulence is much less understood. Identifying and characterizing small-molecule virulence factors reveals new virulence pathways that could be targeted to control infections. Genome analyses of pathogenic bacteria have uncovered many gene clusters for the biosynthesis of small molecules; the challenge lies in determining the chemical structures of these molecules. We focus on Pseudomonadaceae, an important family of Gram-negative pathogens that infect both plants and humans. We take a multidisciplinary approach to identify novel small-molecule virulence factors to study the chemical interactions between bacteria and their hosts.
3. Deciphering the biosynthesis of natural-product antibiotics
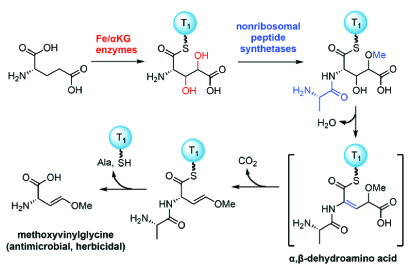 Understanding how bacteria synthesize natural products is essential for identification and engineering of natural products. To augment the discovery and mechanistic studies of natural products, we identify enzymes responsible for their biosynthesis and elucidate the chemistry catalyzed by these enzymes. For example, we study the biosynthesis pathways of natural products comprising nonproteinogenic amino acids (NAAs). NAAs are ubiquitous in bacteria, plants and animals and exhibit antimetabolite, antimicrobial, and agrichemical properties. We characterized the biosynthesis of a unique family of NAAs, the oxyvinylglycines. Our work discovered new enzyme-catalyzed chemistry and generated tools for identifying and engineering of NAAs.
Understanding how bacteria synthesize natural products is essential for identification and engineering of natural products. To augment the discovery and mechanistic studies of natural products, we identify enzymes responsible for their biosynthesis and elucidate the chemistry catalyzed by these enzymes. For example, we study the biosynthesis pathways of natural products comprising nonproteinogenic amino acids (NAAs). NAAs are ubiquitous in bacteria, plants and animals and exhibit antimetabolite, antimicrobial, and agrichemical properties. We characterized the biosynthesis of a unique family of NAAs, the oxyvinylglycines. Our work discovered new enzyme-catalyzed chemistry and generated tools for identifying and engineering of NAAs.
